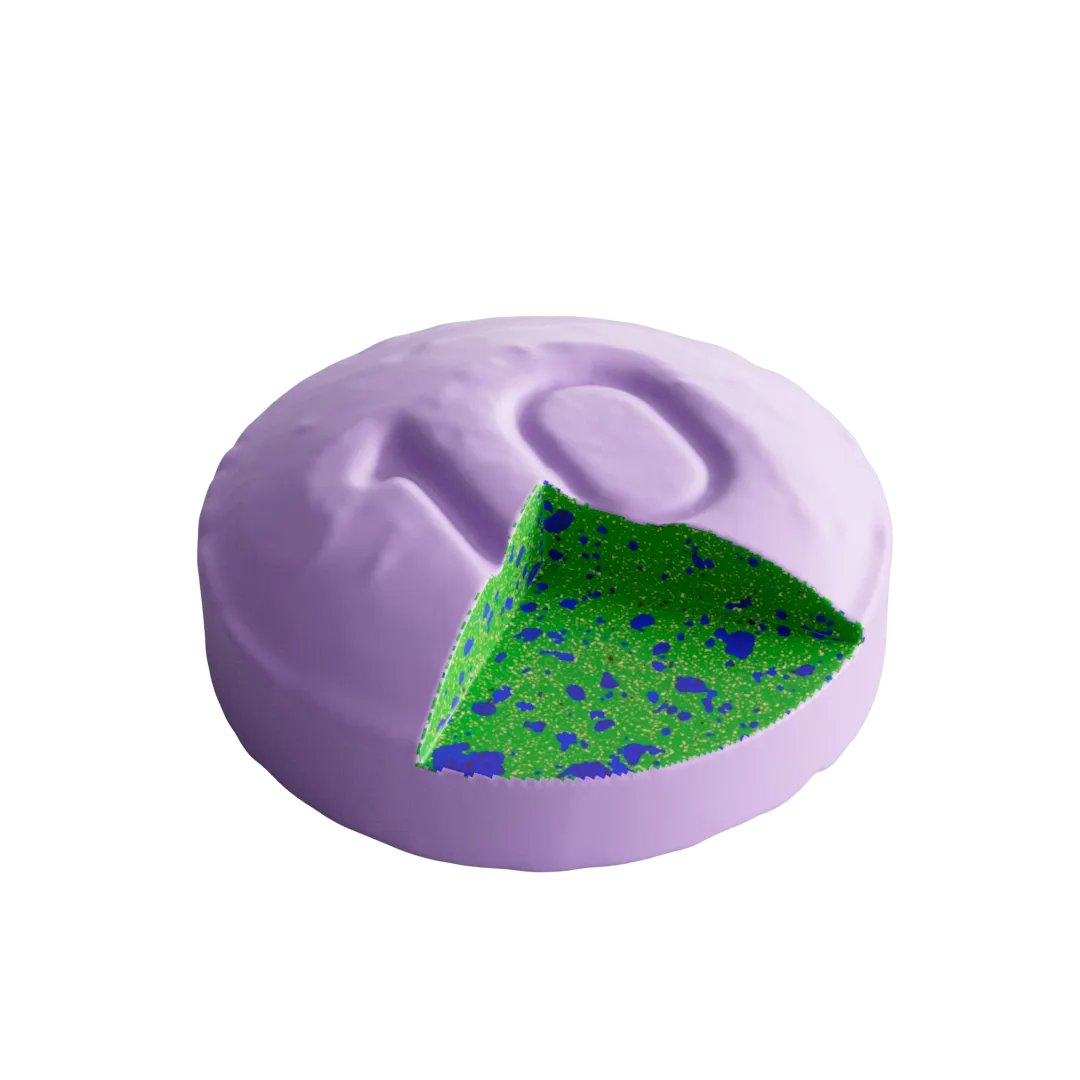
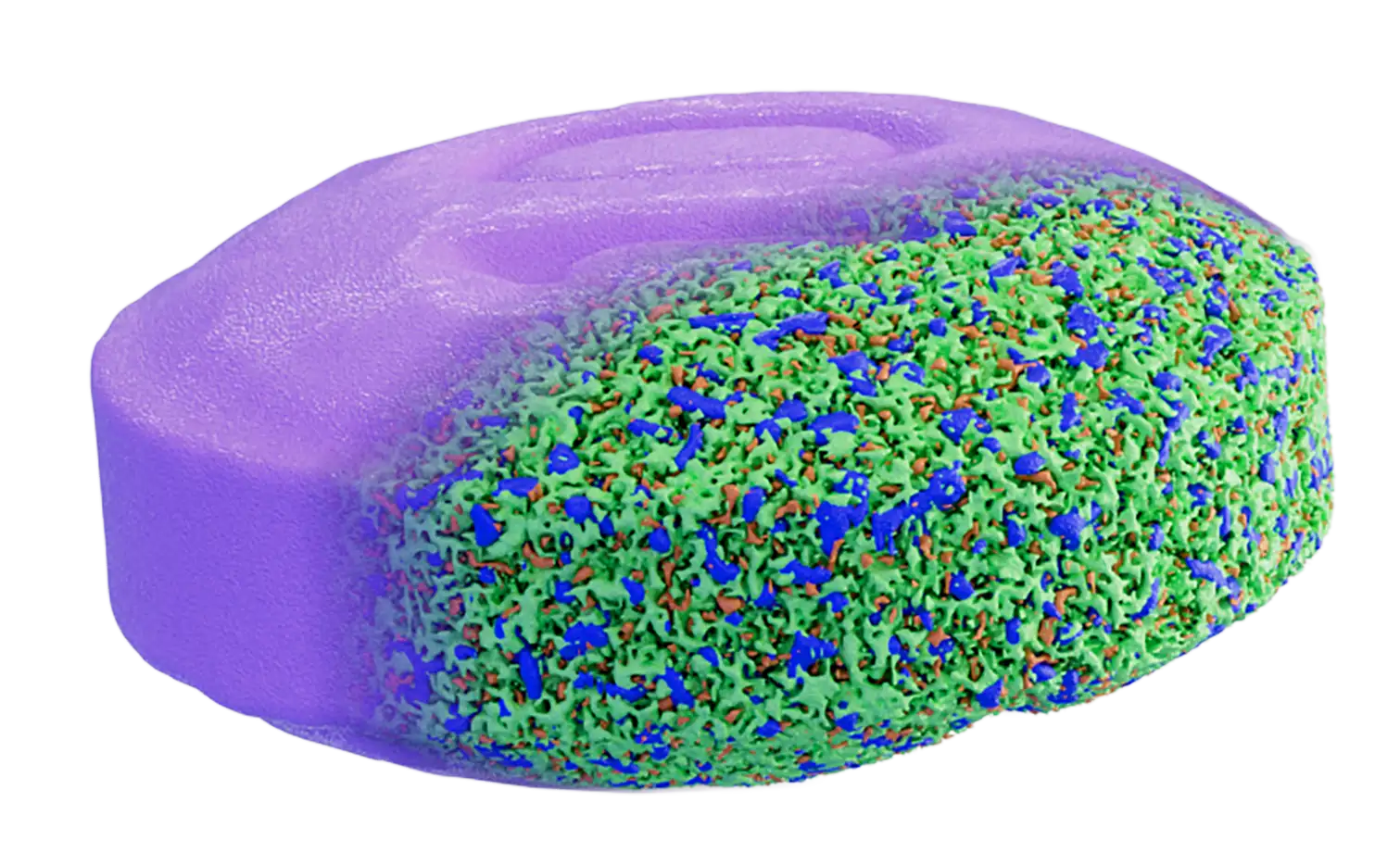
DigiM I2S Software
I2S, or image to simulation, is a cloud based image processing platform designed to manage the complete lifecycle of your image data.

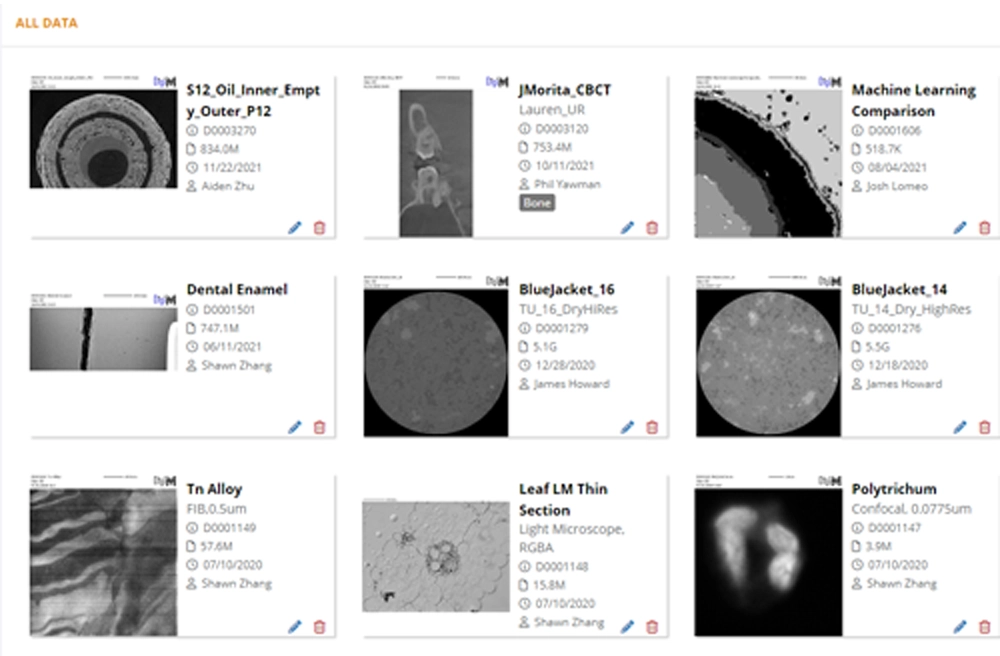
Cloud-Based Data Management
Utilizing a cloud based data management library, I2S allows its users to access image data anytime, anywhere, from any device. I2S data management includes FDA CFR 21 Part 11 compliance, by storing and maintaining these records I2S will automatically and track all data created, modified, and retrieved from within the platform for the data's entire lifecycle.
Machine Learning and Deep Learning
I2S utilizes artificial intelligence based techniques to support the digital transformation of your drug products. Through supervised machine learning and deep learning semantic image segmentation techniques, I2S offers the tools to handle any type of image analysis problem.

Quantifications
Once segmentation has been performed to create a digital twin of a drug product, an assortment of quality attributes can be computed from the twin. Morphological characterizations can be performed to determine metrics such as volume, particle size distribution, and spatial distributions, and many more. Utilizing these measured attributes users can begin to correlate microstructural critical quality attributes to process parameters and product performance in order to gain a greater understanding of their products.

Physics-based Simulations
I2S, features a suite of physics based simulations allowing a user to compute mass transport properties of there product directly from their imagery. digiM has developed a voxel based simulation method, in which each pixel of the scene is used as a computational cells to assist in these computations. This voxel based approach allows I2S users to compute highly accurate mass transport properties much faster than traditional fluid dynamics simulation techniques. I2S also contains a patented diffusion simulation allowing users to predict the drug release of the digital twins of their products.

Begin your data transformation journey
Contact our specialists to get started with a trial















